8. Genome comparison dock¶
The Genome Comparison Dock allows genomes to be inspected and compared within RevoSim, and can be launched from the main menu or the toolbar. By default it appears landscape on the bottom of the main window. Its landscape orientation is intended to allow whole genomes to be viewed without scrolling on most operating systems, however we note that on operating systems with low resolution displays, the REvoSim GUI may struggle to accommodate the docker in its default position: the environment and population views may be forced to be small, or the genome comparison docker may not have the vertical extent to show more than one genome. If this is the case, the docker can be floated, or alternatively docked to the right or left of the main window.
Video 8.1 - Genome comparison dock in use.
A genome can be added to the comparison dock by right clicking on the population scene. This will add the genome of the first digital organism living within that pixel to the comparison dock (i.e. that held in slot zero), rendering it as zeros and ones with coding genome on the left, as well as displaying the colour it is rendered as in genome views, and the colour of its environment. Multiple genomes can be selected and inspected in this manner, and a comparison of two can be achieved by selecting the desired genomes and selecting compare, which renders the two in a lower panel highlighting the differences between the genomes. By default, auto compare is enabled: this compares and highlights the differences between each entry on the genome list and that before it. See Video 8.1 for example of use.
The genomes can be deleted individually from the table by selecting a genome and using the Delete button. Alternatively all genomes in the table can be removed from Genome Comparison Dock the using the Reset All button (this will also remove any manually compared genomes).
The first ten genomes from the Genome Comparison Dock are pulled through to the reseed dialogue (Fig. 8.1): this allows one to be selected as the starting genome for a new run. This allows multiple repeats of simulations using the same genome, which is known to be capable of surviving within a given environment using the masks loaded at launch (note that these will not persist between sessions unless through loading a saved simulation).
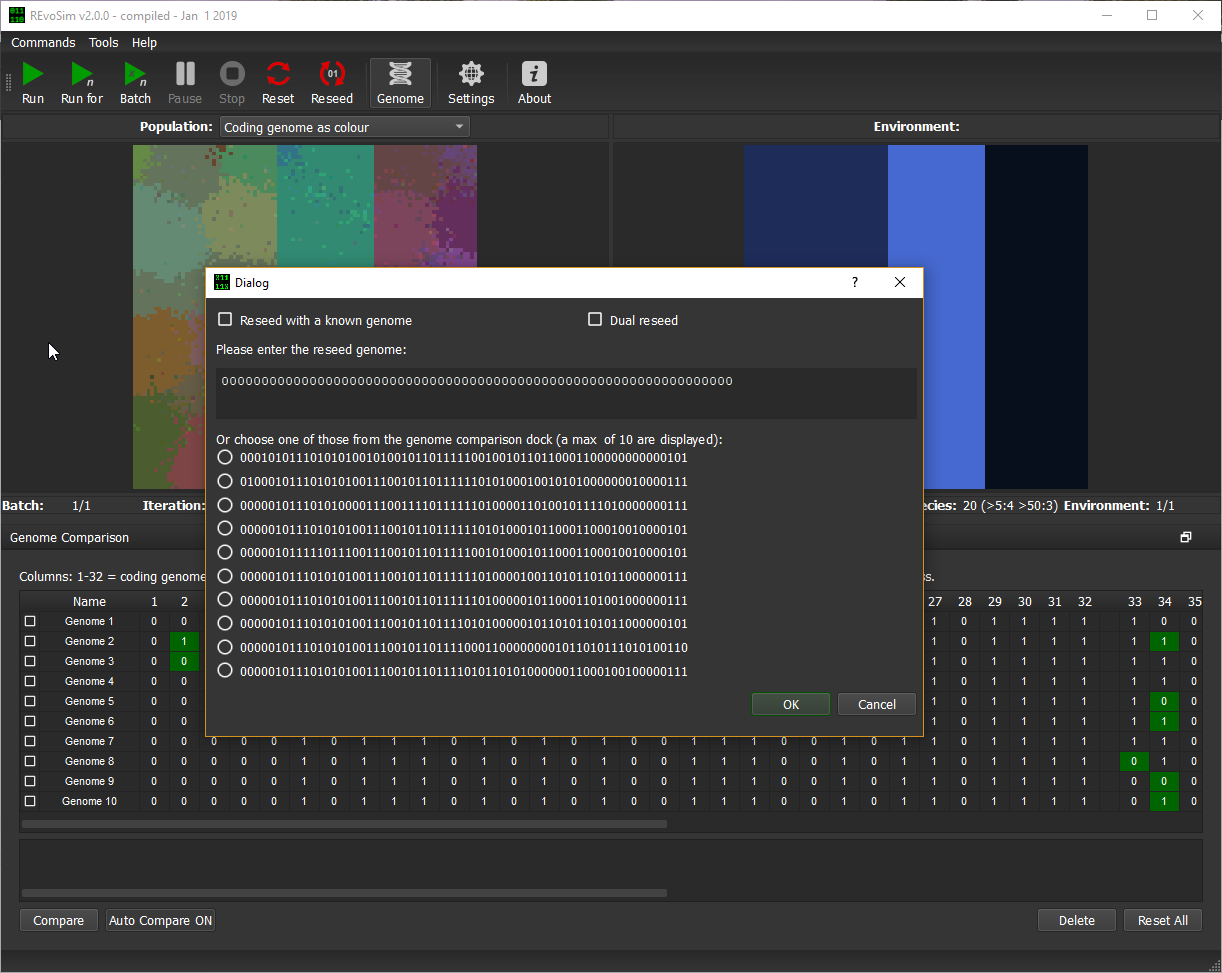
Figure 8.1 - Reseed dialog showing first ten genomes as selected in the Genome Comparison Dock.